In this case study we hear from Ellen Jenkins, a postdoctoral student from the Institute of Inflammation and Ageing. Ellen is looking into developing a deep learning model to automatically differentiate the tissue from the air space.


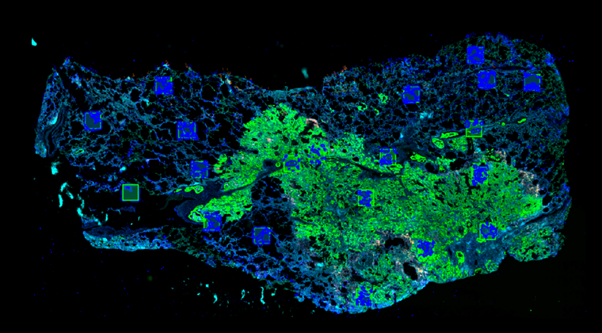
There is an ongoing project in the respiratory research group (CMH, IIA) which utilises samples of human lung tissue that are donated by patients who are receiving a surgery for different lung conditions (cancer, fibrosis, emphysema). Using a wide range of antibodies, different markers in the tissue can be labelled different colours – producing a complex image of cells (figure 1). The lung is divided into two main regions: the air spaces and the supporting tissue. Cells that reside in each of these regions may look identical using this imaging method, however they are thought to play very different roles across different types of lung disease. It is therefore important to separate cells in these images into these two populations dictated by their spatial niche. Manual annotation of these two regions across whole slides is time consuming and not feasible as this project expands in sample number. Therefore, we aimed to develop a deep learning model to automatically differentiate the tissue from the air space.
…increasing the efficiency of my code (reducing run time from 7 hours to a few minutes) by testing and editing using Baskerville Tier 2 HPC
I have completed this workflow previously within commercial software Visiopharm which is licensed for use at UoB by Birmingham Tissue Analytics. This project produced a promising result, demonstrating that segmentation of these images is feasible despite the small number of samples available. The next steps for this project were to design a model that would be transferable to other research groups staining similar tissue, for whom Visiopharm may not be an available resource.
I therefore began to train a u-net style model using PyTorch via BlueBEAR HPC with GPU access. This was with the aim of developing an open-source model that could be accessed by other researchers and fine-tuned via transfer learning to their specific datasets. I have limited experience with python and HPC so received support from the Advanced Research Computing team who corrected my errors, increasing the efficiency of my code (reducing run time from 7 hours to a few minutes) by testing and editing using Baskerville Tier 2 HPC. I was advised about some pitfalls in my current approach and additions that could improve performance of my model, which I applied back to my project on BlueBEAR HPC.
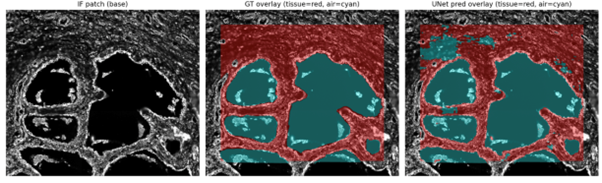
The pipeline now produces accurate overlays compared to ground truth annotation
The pipeline now produces accurate overlays compared to ground truth annotation shown in figure 2. The cells within each label can be seen clearly, highlighting the different populations that can now be examined individually using standard immunofluorescence image analysis pipelines. The model was also tested against an open-source dataset of lung images (Temko-Maliga CyCIF lung adenocarcinoma, PMID: 36419822). Whilst performance was not completely accurate, the model made sensible predictions and was subject to the same failure modes observed within my original dataset. This suggests it could be a promising platform for transfer learning, which we aim to publish as a methods manuscript by July 2026.
This work was supported by The Kennedy Trust for Rheumatology Research [grant no. KENN 20 21 04]
We were so pleased to hear of how Ellen was able to make use of what is on offer from Advanced Research Computing, particularly to hear of how they have made use of the Baskerville compute – if you have any examples of how it has helped your research then do get in contact with us at bearinfo@contacts.bham.ac.uk.
We are always looking for good examples of use of High Performance Computing to nominate for HPC Wire Awards – see our recent winner for more details.
